"If you have a damaged cell, you don't want it to replicate," explained computational biophysicist Rommie Amaro. She and her team have been modeling the tumor suppresion protein called p53 using the Stampede supercomputer at the Texas Advanced Computing Center - a virtual tour is shown in the video below. She has been at work on this molecule for many years, especially driven by the fact that the p53 protein can prevent the formation of malignant cells. p53 is sometimes called the guardian of the genome because it is so important as a regulator of the life cycle of a cell.
Her new work is based on the full length version of the p53 gene, which presents challenges. It has a complex structure and several regions with a high degree of flexibility. "We went from studying a relatively small protein of 50,000 atoms in a water box to the full length p53, which included the single binding domain, additional domains, in tetramer form, and in complex with different segments of DNA," Amaro explained. "In doing so, there was a huge increase in terms of the complexity of the system and the number of atoms simulated."
"That's why Stampede was so terrific," Amaro continued. "Stampede has many processors and amazing scalability -- we were able to run this 1.5 million atom system for nearly a microsecond and actually begin to say things about the dynamics at a completely different scale than what was known previously.
Amaro thinks of these techniques as a sort of computational microscope. “The science challenge represents a level of complexity that's very difficult if not impossible to experimentally test," she said. In response, the researchers built atomic-level models 'in silico,' and interrogated the system in unique ways. For the first time, for example, the researchers built a large complex of the p53 molecule with three different sequences of DNA: two were recognition sequences (P21 and Puma) and the third strand was a negative control.
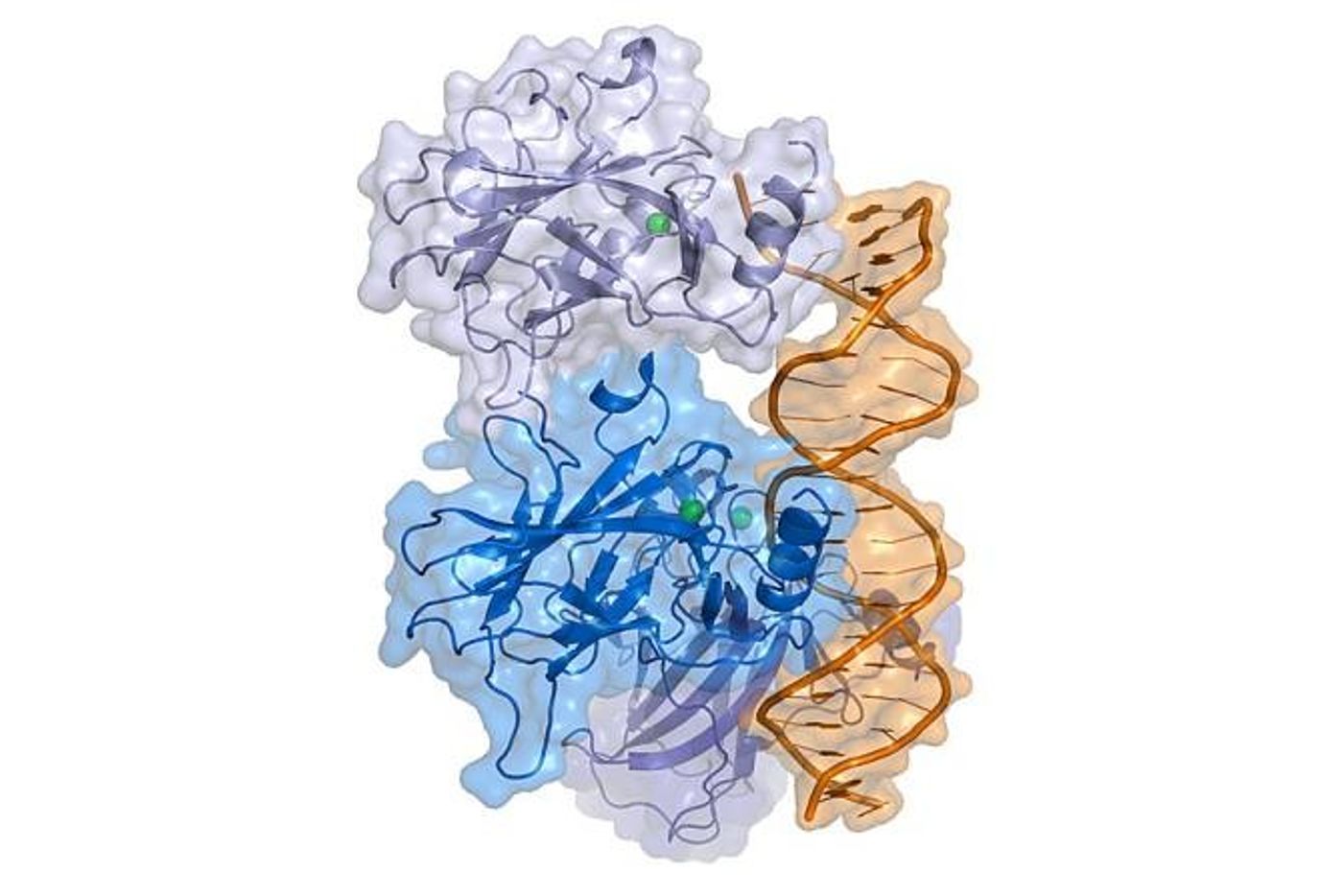
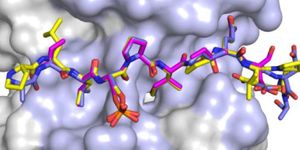
This was the first time the p53 molecule was seen directly interacting with DNA, and the first time a change in that interaction was seen - a change dependent on the sequence of the DNA. "We could see how when we the full-length p53 was bound to a DNA sequence that was a recognition sequence, the tetramer clamps down and grips onto the DNA - which was unexpected," Amaro said. In contrast, with the negative control DNA p53 stays more open. "It actually relaxes," she noted. "It suggested a mechanism by which this molecule could actually change its dynamics depending on the exact sequence of DNA."
"We have all of the atoms and the proteins," Amaro continued. "We try to replicate the environment of the cell, so we have water molecules and ions, and then we had an actual segment of DNA so we could explore the response of 'full length p53' in terms of its dynamics, depending on what sequence of the DNA was there."
The researchers used three different systems and produced three different versions of each system to check for variability in the data; their strategy created a total of nine different simulations for nearly a microsecond of aggregated dynamics. "The simulations were very computationally intensive. And then to be able to do something new about the biology that wasn't known before - that's the really exciting part and that's what we showed," Amaro said.
"As the systems get bigger, much more computational time is required. Previously the simulations were run for a few nanoseconds. Now we have microseconds of dynamical data, which is 1,000x more. In this case we were looking at over a dozen domains and in complex with the DNA. It gives us a much more complete picture of what is actually happening. It's a few steps closer to reality than anything we've been able to accomplish yet," she commented.
"Computing is getting to the point now where it can have an impact on developing new therapies. It gives us a better understanding of cancer mechanisms and ways to develop possible novel therapeutic avenues. When most people think about cancer research they probably don't think about computers, but these models are getting to the point where they have a great impact on the science." Amaro wants to see these discoveries used to create new cancer therapeutics.
"The ideal situation is that cancers of the breast and prostate could possibly be rescued or eliminated if we were able to develop a compound that reactivated p53. You don't have to wait until the cancer is advanced. The sensing of mutation and then shuttling the cell to die is something that happens through the early stages of cancer. It could be detected and fixed before the cancer develops," concluded Amaro. There is more about her work in the video above.
Sources:
AAAS/Eurekalert! via
Texas Advanced Computing Center,
Oncogene