Artificial Intelligence Provides Cancer Insights
Artificial intelligence has helped researchers learn more about the biophysics of cancer. Research teams that worked in collaboration at Tufts University’s School of Arts and Sciences, the Allen Discovery Center at Tufts, and the University of Maryland, Baltimore County developed a machine-learning platform. It was used to predict a trio of reagents that was would cause symptoms resembling cancer in tadpoles. In the video below hear about the work, which was published in the journal Scientific Reports, from a lead scientist.
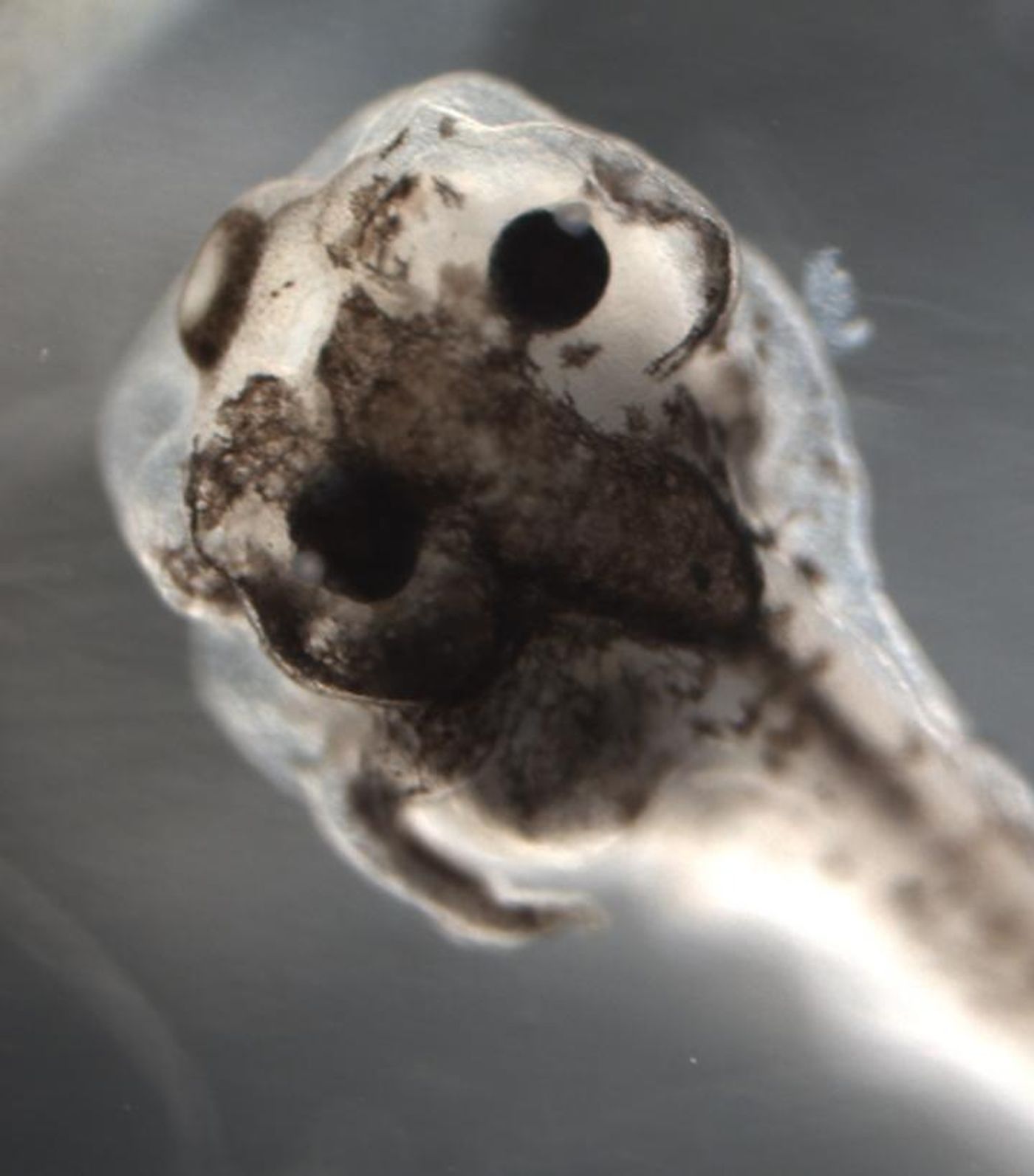
Previous studies by the scientists have indicated that pigment cells, melanocytes, of developing frogs became metastatic, or cancer-like, after normal bioelectric and serotonergic signaling was perturbed. The investigators also utilized AI in to reverse engineer a model of this complicated system. The researchers took note of something that happened throughout their work - the melanocytes of individual frog larva either became cancer-like, or they stayed totally normal.
The researchers were intrigued by this all-or-nothing commitment by the melanocytes. In their latest work, they have used the model they made with AI to try to achieve only a partial change of the melanocytes, in an effort to learn more about cell behavior.
"We wanted to see if we could break the concordance among cells, which would help us understand how cells make group decisions and determine complex body-wide outcomes," explained the corresponding author of the study, Michael Levin, Ph.D., Vannevar Bush Professor of Biology and Director of the Allen Discovery Center at Tufts and the Tufts Center for Regenerative and Developmental Biology.
AI opted for a specific recipe of three ingredients - altanserin, a 5HTR2 inhibitor; reserpine, a VMAT inhibitor, and VP16-XlCreb1, mRNA encoding constitutively active CREB - to meet the partial-conversion goal. Exposing live tadpoles to the special cocktail did indeed achieve something that hadn’t been seen before, partial melanocyte conversion within single frog larvae.
"Our system predicted a three-component treatment, which we'd never have come up with on our own, that achieved the exact outcome we wanted, and which we hadn't seen before in years of diverse experiments. Such approaches are a key step for regenerative medicine, where a major obstacle is the fact that it is usually very hard to know how to manipulate the complex networks discovered by bioinformatics and wet lab experiments in such a way as to reach a desired therapeutic outcome," said Levin.
"Much of biomedicine boils down to this: We have a complex biological system, and a ton of data on what various perturbations have been seen to do to it. Now we want to do something different--cure a disease, control cell behavior, regenerate tissue. For almost any problem where a lot of data are available, we can use this model-discovery platform to find a model and then interrogate it to see what we have to do to achieve result X," added Levin.
It took the AI-discovered model 576 virtual experiments, which computationally modeled the development of an embryo under a different novel combination of drugs 100 times, to eventually identify the perfect combination.
"Even with the full model describing the exact mechanism that controls the system, a human scientist alone would not have been able to find the exact combination of drugs that would result in the desired outcome. This provides proof-of-concept of how an artificial intelligence system can help us find the exact interventions necessary to obtain a specific result," said the first author of the report, Daniel Lobo, Ph.D., who worked in the Levin laboratory and is now Assistant Professor of Biology and Computer Science at the University of Maryland, Baltimore County.
Colaborating with Levin and Lobo on the study was Maria Lobikin, Ph.D., a former Levin laboratory member who is currently a scientist at Homology Medicines Inc.
Future work aims to broaden use of the model and include time-series data, hopefully to provide even more accurate computer models of live systems. Additionally, researchers want to apply this methodology to other improve tumor reprogramming, boost cell regeneration and gain fine control of stem cells.
Sources: AAAS/Eurekalert! via Tufts University, Scientific Reports