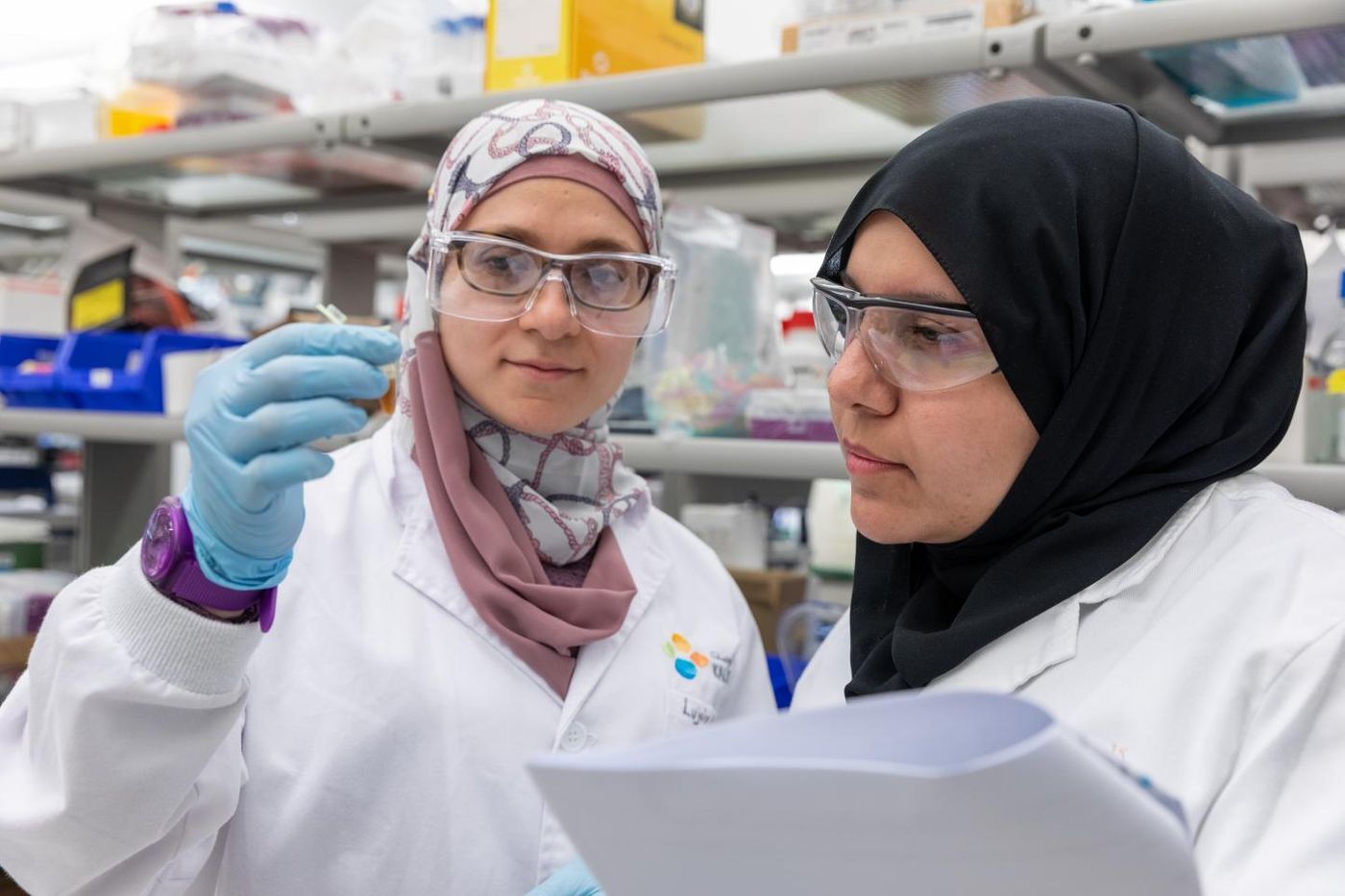
Finding Cancer-Promoting Genes Using Machine Learning
Machine learning algorithms are increasingly being applied to the vast amount of genetic data that has been generated over recent decades. Scientists have now developed a method to identify the genes that encourage tumors to grow, by using computational tools to assess data from mouse models, cancer cell lines, and patients. Genes with causative and potentially causative roles were found for twenty kinds of tumors. The technique, which uses artificial intelligence to connect DNA mutations to functional changes, is available for free online and has been published in Scientific Reports. It may help advance personalized medicine or drug development.
Algorithm turns cancer gene discovery on its head from KAUST Discovery on Vimeo.
"Our method can be used as a framework to predict and validate cancer-driver genes in any database or real population sample," said first study author Sara Althubaiti, a graduate student in the lab of research leader Robert Hoehndorf of KAUST's Computational Bioscience Research Center.
It has been laborious and time-consuming to find genes that play a role in the development of cancer. Tumor samples have to be gathered from patients and the genetic information from those cancer cells has to be sequenced, then compared to other samples from cancer patients and healthy individuals. The findings from these investigations are then typically studied in models like cell culture or rodents.
"Our method turns this approach on its head," said Althubaiti. "Essentially, our approach is knowledge-driven and we use tumor sequencing data as validation. This is unlike most approaches, which are data-driven combined with interpretation of the findings with respect to established knowledge."
The KAUST team is aiming to find new genes that are involved in cancer; such discoveries have been in decline in recent years. This work takes biological pathways and characteristics that are related to tumor growth into account. The algorithm is meant to recognize patterns in the functions of genes that make them more likely to drive tumor development. The researchers applied their tool to a publicly available database containing tumor variants, and showed that the algorithm was able to find genes that promote cancer and other likely candidates.
The scientists also used the algorithm to assess samples from people with a rare cancer, nasopharyngeal carcinoma, and individuals with colorectal cancer. The computational tool identified potential cancer-driving genes that were often mutated and had features in common with other cancer-causing genes.
"This work is a good example for scientific collaboration within Saudi Arabia," Hoehndorf said, "but it also demonstrates the need for multidisciplinary collaborations between computer scientists, clinical researchers and biologists."
Sources: AAAS/Eurekalert! via King Abdullah University of Science & Technology (KAUST), Scientific Reports