A modular system of proteins can detect a particular DNA sequence in a cell and then trigger a specific response, such as cell death, according to biological engineers at the
Massachusetts Institute of Technology (MIT). They can customize the system to detect any DNA sequence in a mammalian cell and then launch a desired response, including killing cancerous or virus-infected cells.
According to James Collins, the Termeer Professor of Medical Engineering and Science in MIT’s Department of Biological Engineering and Institute of Medical Engineering and Science (IMES), “There is a range of applications for which this could be important. This allows you to readily design constructs that enable a programmed cell to both detect DNA and act on that detection, with a report system and/or a respond system.”
Collins is the senior author of a Nature Methods paper describing the technology, which was reported in
Drug Discovery & Development. The technology is based on DNA-binding proteins called zinc fingers. The proteins can be designed to recognize any DNA sequence.
Shimyn Slomovic, an IMES postdoc and the paper’s lead author, added, “The technologies are out there to engineer proteins to bind to virtually any DNA sequence that you want. This is used in many ways, but not so much for detection. We felt that there was a lot of potential in harnessing this designable DNA-binding technology for detection.”
The researchers linked the DNA-binding capability of zinc fingers with a consequence of either turning on a fluorescent protein to reveal that the target DNA is present or generating another type of action inside the cell. They exploited a short protein called intein that can be inserted into a larger protein, splitting it into two pieces. The split protein pieces, exteins, become functional when the intein removed itself while rejoining the two halves.
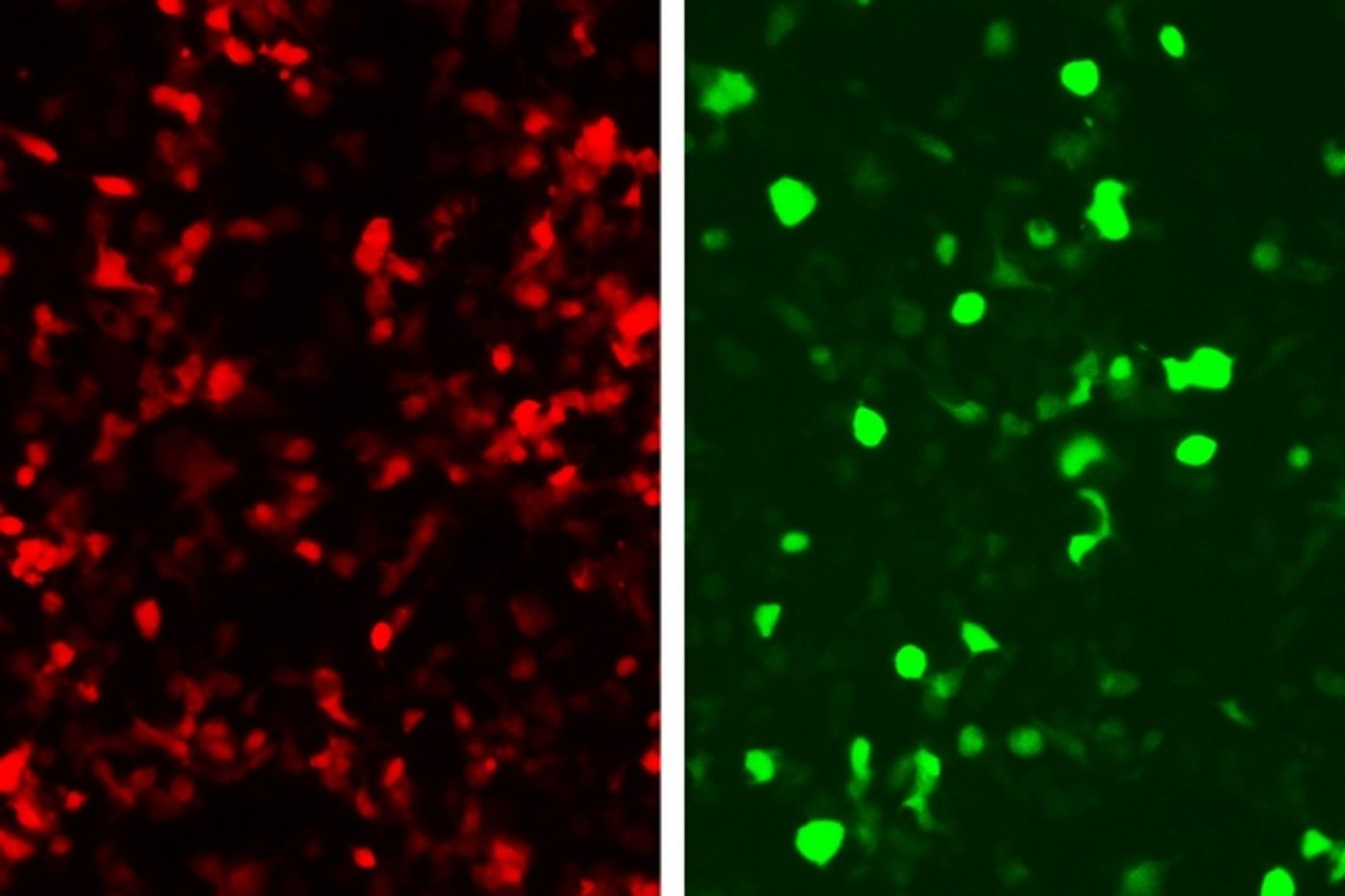
The researchers then divided an intein in two and attached each portion to a split extein half and a zinc finger protein. They engineered the zinc finger proteins to recognize adjacent DNA sequences within the targeted gene. If they both find their sequences, the inteins line up and are cut out, enabling the extein halves to rejoin and form a functional protein. The researchers explained that the extein protein is a transcription factor that can turn on any chosen gene. They linked green fluorescent protein (GFP) production to the zinc fingers’ recognition of a DNA sequence from an adenovirus, enabling any cell infected with this virus to glow green.
This approach could be used both to reveal infected cells and to kill them by programming the system to produce proteins that alert immune cells to fight the infection, instead of GFP. As Slomovic said, “Since this is modular, you can potentially evoke any response that you want. You could program the cell to kill itself, or to secrete proteins that would allow the immune system to identify it as an enemy cell so the immune system would take care of it.”