A Supercomputer Aids in the Hunt for COVID-19 Therapeutics
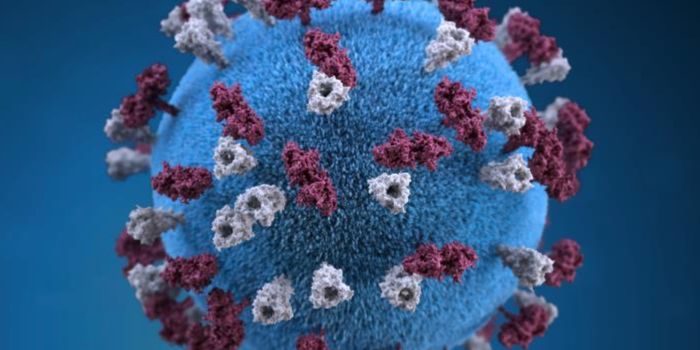
It is seeming more likely that COVID-19, caused by the coronavirus SARS-CoV-2, will impact many people. While an antiviral drug called remdesivir is being tested as a therapeutic for the illness in clinical trials, a different group of scientists at Oak Ridge National Laboratory (ORNL) is using a supercomputer to find small molecules that might be effective at battling the viral infection. The work, reported on ChemRxiv, used the most powerful supercomputer in the world, Summit, to screen 8,000 compounds and reveal 77 candidates that are most likely to attach to the main protein on the SARS-CoV-2 virus and prevent it from infecting cells.
We know how the coronavirus SARS gets into cells, and when researchers in China sequenced the genome of SARS-CoV-2, it was suspected that it uses a similar mechanism to enter a host.
"We were able to design a thorough computational model based on information that has only recently been published in the literature on this virus," said ORNL CMB postdoctoral researcher Micholas Smith, who referenced a recently published study in Science China Life Sciences. Smith composed a model of the viral spike protein, called the S-protein. Spike proteins bind to receptors on host cells, and enable them to gain entry.
Smith used the computational time he was granted on Summit to utilize a chemical simulations code, and perform molecular dynamics simulations. These simulations model the movements of a protein's atoms and particles. This analysis enabled him to simulate how various compounds would bind to the coronavirus S-protein, and whether any stopped it from linking up with human cells.
"Using Summit, we ranked these compounds based on a set of criteria related to how likely they were to bind to the S-protein spike," Smith said. A more accurate version of the S-protein model has now been reported in Science, and the team is planning to repeat their computational work with this updated version. It may change their list of 77 candidates. After that, any potential compounds will have to be tested experimentally for efficacy.
"Summit was needed to rapidly get the simulation results we needed. It took us a day or two whereas it would have taken months on a normal computer," said study author Jeremy Smith. "Our results don't mean that we have found a cure or treatment for the Wuhan coronavirus. We are very hopeful, though, that our computational findings will both inform future studies and provide a framework that experimentalists will use to further investigate these compounds. Only then will we know whether any of them exhibit the characteristics needed to mitigate this virus."
Sources: Phys.org via ORNL, ChemRxiv