Drug Resistance in Tuberculosis Involves a Unique Mechanism
The pathogenic bacterium that causes tuberculosis, Mycobacterium tuberculosis, does not multiply quickly, so researchers have long wondered how it is able to develop drug resistance as rapidly as it does; it often only takes a few weeks or months to do so. New research reported in eLife has suggested that the process does not involve a piece of DNA that microbes can share (a plasmid or mobile genetic element) or a change in its genome. Instead, it may be due to epigenetic changes.
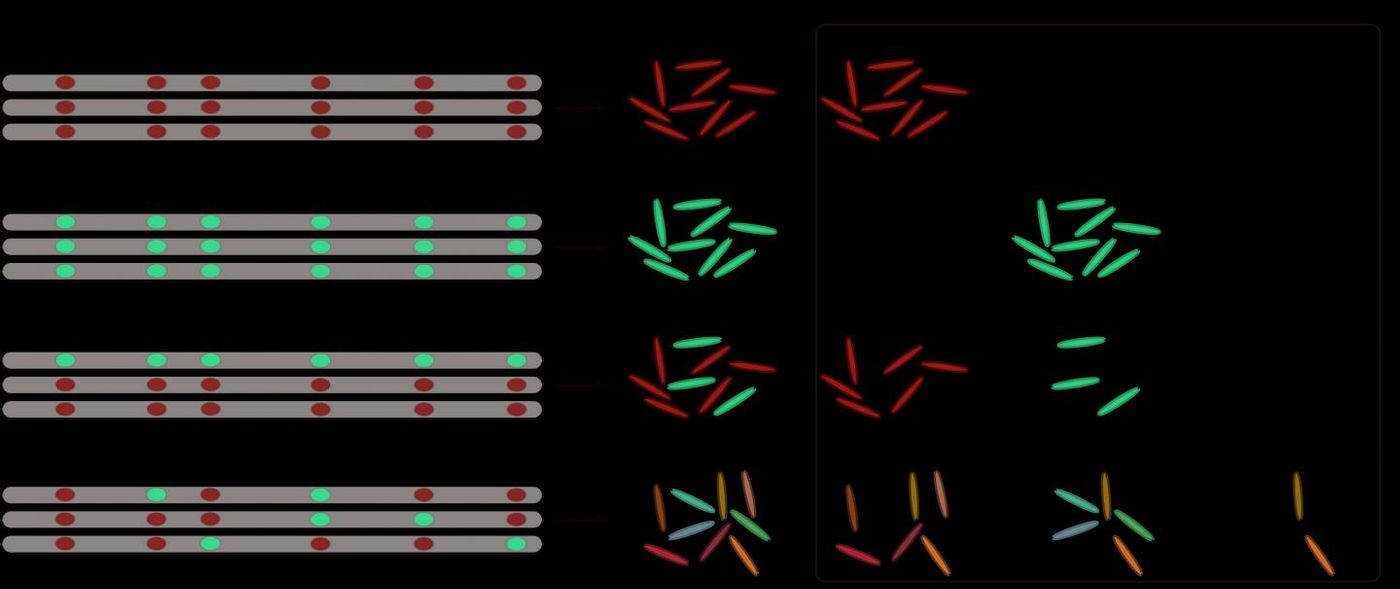
Epigenetics refers to biochemical changes that occur on the genome that affect gene expression, and involve chemical tags like methyl groups that are added or structural alterations. These changes don't affect the DNA sequence, but some have been shown to be passed down to future generations. If epigenetics is the key to drug resistance in M. tuberculosis, researchers may be able to develop better treatments or vaccines for TB.
This new work suggests that M. tuberculosis can rapidly produce multiple subgroups that have their own physical characteristics or phenotypes. Though antibiotics are able to kill most of these populations off, a few will survive, and these microbes will be drug-resistant.
"We believe this also explains why diagnostic testing in some patients does not predict treatment failure, and why some patients come back months later with the disease reemerging in a far more resistant state," said Faramarz Valafar, a TB expert with San Diego State University's School of Public Health. "This is also why CT scans of the lungs of many "cured" patients show lesions with possible bacterial activity."
For this study, the researchers collected hundreds of samples from patients and research labs around the world infected with TB, many of which are drug-resistant.
TB killed an estimated 1.4 million people in 2019 and the pathogen sickens around ten million people every year, according to the World Health Organization.
"We've known for decades that bacterial epigenetics can influence the expression of certain genes, which can lead to a variety of phenotypes even when they have identical genotypes," said graduate student Derek Conkle-Gutierrez. "We discovered evidence of that phenomenon in the TB bacterium."
"We found that some of them had mutations that led to variable DNA methylation and those strains had much more diversity in their epigenome, and thus more potential to be drug-resistant," said project scientist Samuel Modlin.
The study did not reveal any patterns, but it involved an epigenetic feature called methylation. Future work will focus on the genes that are being affected by these methyl groups, and eventually, it could be applied to the development of treatments or diagnostics.
"There is a lot of resistance in TB that escapes current molecular diagnostics and we don't really know why. That's problematic," Valafar said. "This study offers a new domain, new tools, and a new approach to looking for alternative mechanisms. We move away from the classical view of molecular diagnostics and use a novel, comprehensive approach to analyzing bacteria."
Sources: AAAS/Eurekalert! via San Diego State University, eLife