Using a special type of microscopy, researchers were able to capture changes in the nucleus of nerve cells as they underwent differentiation – a process through which they become more specialized cells with specific functions.
Published in Cell Reports, this data has informed the understanding of cellular architecture and may inform research into how genes are regulated.
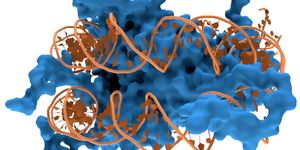
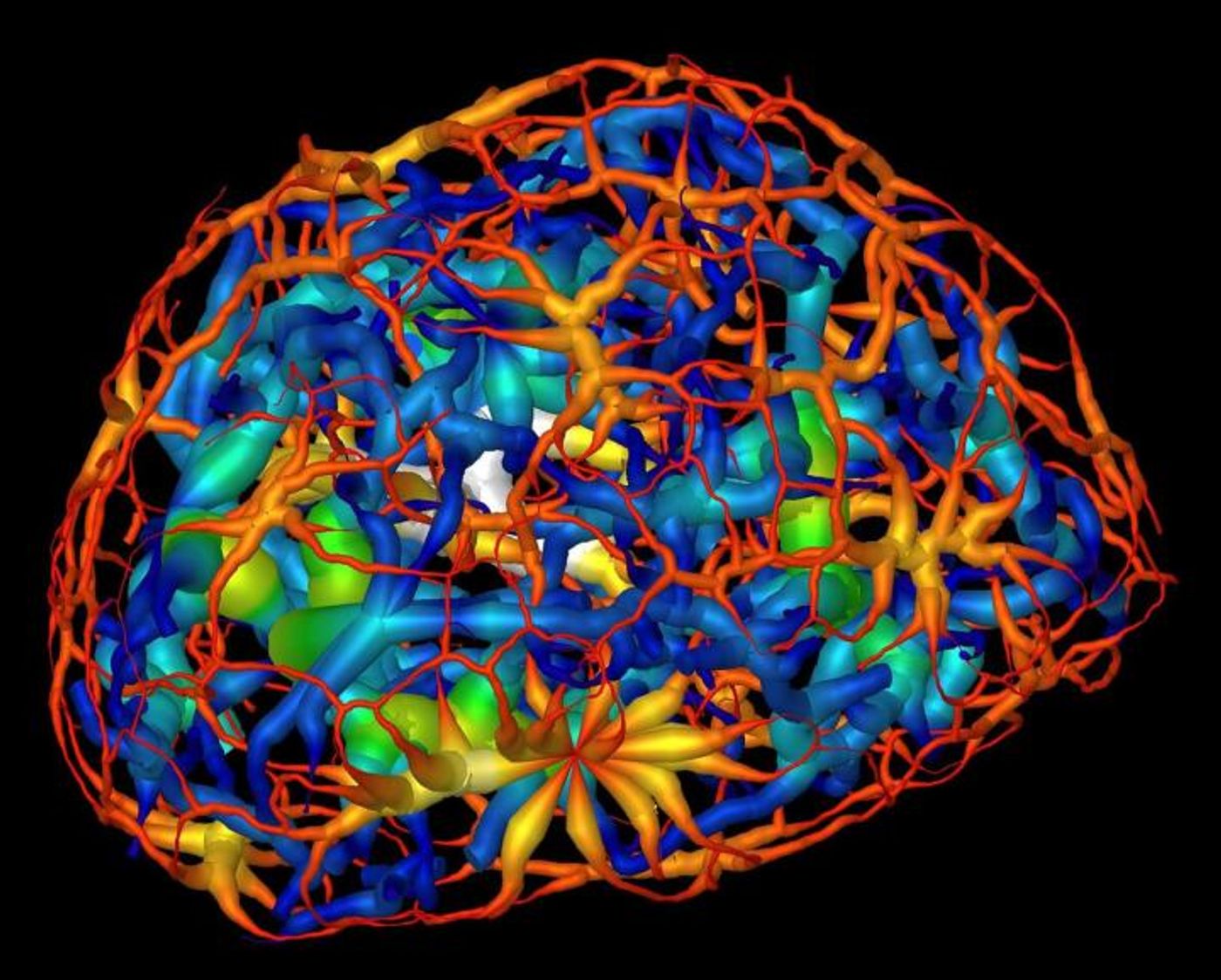
Researchers used powerful X-ray microscope at Berkeley Lab's Advanced Light Source (ALS) to study differentiating neurons as they mature. Using X-rays, images were taken from many angles to create a three-dimensional composite image from two-dimensional images, showing the changing structure of chromatin during the maturation of olfactory neurons of mice. Chromatin functions to package DNA so that it can fit inside of a cell, and forms chromosomes. In the video above, take a trip through the nucleus of a cell; in the video below, DNA reorganization in the nucleus a mouse cell is illustrated.
The investigators were also able to take measurements of the packing inside of a type of chromatin, heterochromatin, and revealed more about the role of a protein that aids in the packaging of heterochromatin, and its localization to the nucleus.
"It's a new way of looking at the nucleus where we don't have to chemically treat the cell," explained Carolyn Larabell, the Director of the National Center for X-ray Tomography (NCXT), a collaboration of Berkeley Lab and UC San Francisco (UCSF). "Being able to directly image and quantify changes in the nucleus is enormously important and has been on cell biologists' wish list for many years."
She added that Chromatin is "notoriously sensitive" to chemical stains and additives frequently used in biological imaging protocols that highlight regions of interest in a sample. "Until now, it has only been possible to image the nucleus indirectly by staining it, in which case the researcher has to take a leap of faith that the stain was evenly distributed."
Larabell, who is also a faculty scientist at Berkeley Lab and a UCSF professor, explained that it had been thought that chromatin existed as a group of disconnected islands, but the latest report indicated that chromatin is compartmentalized into two areas of "crowding" that comprise a continuous network throughout the nucleus.
"We were really surprised: There are no islands, it's all connected," she added. "We could see how chromatins pack through the nucleus and how molecules move through the nucleus, and we found that heterochromatin is 30 percent more crowded than the region where active genes are. That cannot be done with any other imaging techniques." Two-dimensional images would have shown the nucleus as a "flat, confusing mess."
The scientists also wanted to learn more about the expression of genes related to olfaction; mice have around 1500 genes related to the sense of smell. Every olfactory neuron uses one of those genes to make a receptor that detects a set of related odors. The many receptors of those genes enable mice to smell a huge variety of scents.
"We're trying to understand how the reorganization of chromatin affects gene expression," Larabell continued. "No one's been able to study this at the human level yet." This research could potentially help provide new insights about diseases and disorders in which gene expression is a factor. The results of this study are already being used to inform models of cell development.
"This work highlights the power of multidisciplinary research," commented lead author Mark Le Gros, a physicist who was responsible for the design and construction of the X-ray microscope and Associate Director of the NCXT. "This is an example of work that required a combination of molecular biologists and cell biologists with physicists and computer scientists," concluded Le Gros.
Sources:
AAAS/Eurekalert! via
Berkeley Lab,
Cell Reports