PanCancer Atlas of 33 Cancer Types for Genomic Profile Index Completed by NIH TCGA
PanCancer Atlas
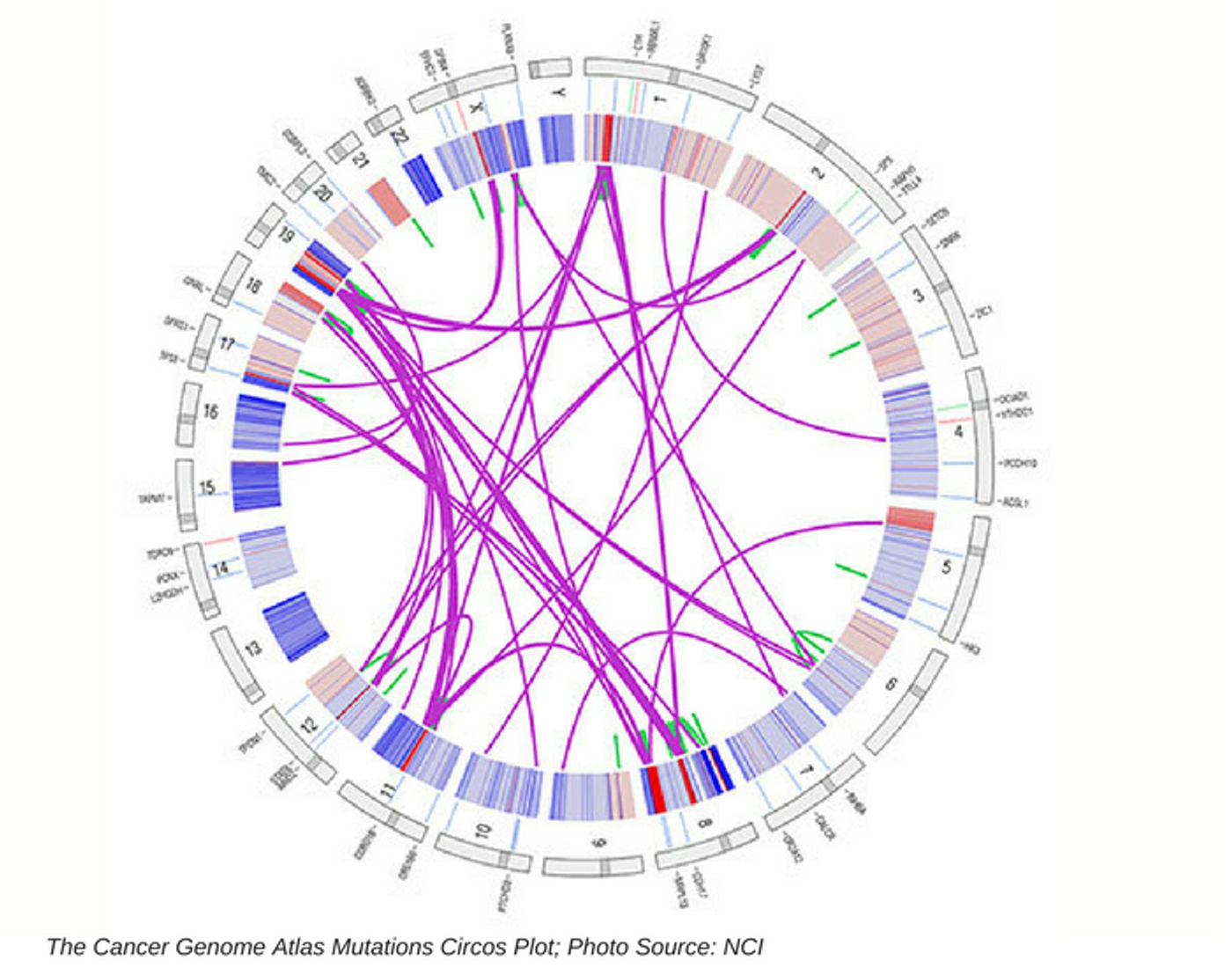
The Cancer Genome Atlas (TCGA) project was created to collect genomic profiles of cancers in one place. The atlas was publically funded by the National Institutes of Health (NIH) and is available to provide better diagnostic and prognostic methods, as well as to prevent cancer where possible. The work was initially launched in 2005 and has been progressing following the advent of technological developments in the fields of genome sequencing and bioinformatics. The demand for better testing, diagnosis, treatment, and prognosis is ever present in healthcare today. The atlas work is truly aimed at creating a database of cancer genomic profiles for numerous cancers or tumors. The NIH TCGA announced their project completion in early April 2018. The PanCancer Atlas is the resulting dataset that incorporates genomics, biological, laboratory science, and health outcome data analyzed using combinations of mathematics, engineering, computer science.
Cancer is complex and more than 200 forms of cancer have been described through their differing profiles, including biological, environmental, and molecular factors. Tomczak, et al., (Contemporary Oncology) wrote of TGCA, “The architecture of occurring genetic aberrations such as somatic mutations, copy number variations, changed gene expression profiles, and different epigenetic alterations, is unique for each type of cancer.” TCGA had a goal to document and create an index of over 30 tumors based on the above listed factors.
Phase I
TGCA was completed in stages and Phase I lasted 3 years. The project needed to develop an analytical framework for incorporated data. Next, they needed to validate their research methods and structure for obtaining and analyzing that information using their developed framework. Phase 1 focused on three tumors with poor prognosis including brain and lung cancers.
Phase II
In 2009, Phase II work began by expanding the analysis to another 30 tumors. This work continued through 2014. There were collaborative supporters and funders of this biomedicine project for the betterment of healthcare and society including institutions across the US and Europe.
Datasets
The PanCancer Atlas characterizes the genomic differences between normal and 33 prevalent cancerous tumor types. Analysis and datasets include information about numerous factors or changes involved in cancer development. The different methods and datasets include:
-
RNA Sequencing – total RNA profiling to identify and quantitate common and less common transcripts, gene fusions, coding and non-coding sequences.
-
MicroRNA sequencing – detection and identification of short, noncoding RNAs which can potentially regulate genes across multiple signaling pathways, and tissue-specific RNA expression.
-
DNA sequencing – Sanger Sequencing method used to provide DNA alteration information such as Copy Number Variants, mutation frequency, insertions, deletions, and polymorphisms.
-
SNP platforms – Array based analysis useful in categorizing structural variation across genomes.
-
Array-Based DNA methylation sequencing – DNA methylation profiles are being found to provide rich information about epigenetic genome alterations affecting gene expression and protein formation.
-
Protein Array – reverse phase analysis which can detect and quantitate miniscule amounts of protein (nanograms) for protein expression profiling, biomarker identification, and diagnostic oncology indicators.
The availability of this comprehensive dataset will be helpful for clinicians and researchers across the globe. With the infrastructure in place, and as more is discovered about the molecular biology of cancer, perhaps additional cancers can be included in this atlas.
Cancers included in the PanCancer Atlas: Breast Ductal Carcinoma, Breast Lobular Carcinoma, Glioblastoma Multiforme, Lower Grade Glioma, Adrenocortical Carcinoma, Papillary Thyroid Carcinoma, Paraganglioma, Pheochromocytoma, Cholangiocarcinoma, Colorectal Adenocarcinoma, Esophageal Cancer, Liver Hepatocellular Carcinoma, Pancreatic Ductal Adenocarcinoma, Stomach Cancer, Cervical Cancer, Ovarian Serous Cystadenocarcinoma, Uterine Carcinoma, Uterine Corpus Endometrial Carcinoma, Head and Neck Squamous Cell Carcinoma, Uveal Melanoma, Acute Myeloid Leukemia, Thymoma, Cutaneous Melanoma, Sarcoma, Lung Adenocarcinoma, Lung Squamous Cell Carcinoma, Mesothelioma, Chromophobe Renal Cell Carcinoma, Clear Cell Kidney Carcinoma, Papillary Kidney Carcinoma, Prostate Adenocarcinoma, Testicular Germ Cell Cancer, and Urothelial Bladder Carcinoma.
Sources: National Institutes of Health, The Cancer Genome Atlas, Contemporary Oncology, Nature.com,