Researchers have created a new, fluorescent probe that detects a specific methyl group on histones in living cells. This tool, a mintbody or modification-specific intracellular antibody, will hopefully lead to a better understanding of histone modification dynamics, how they are regulated, and the impacts on gene regulation and DNA protection.
As seen in the video above, chromatin is a macromolecular complex of RNA, DNA and proteins. It has several important functions, including packaging DNA so it will fit inside of the nucleus of a cell. Chromatin also aids in DNA replication and has an impact on gene regulation. Histones are the part of chromatin that perform the packaging as well as other functions, like DNA repair. Modifications to histone are important to those roles, and you can see more about that in the video below.
One example of such a modification is the addition of a methyl group or groups to histone H4 at the lysine 20 position (H4K20), which is something that is common to many organisms, from yeast to humans. Each methylation state, mono-, di- and trimethylation, each has a unique effect. A major limitation of the study of these changes is that live cells usually have to be fixed at one point in time, and then analyzed. They cannot be kept alive through the process, so the dynamics of histone cannot be observed in the same individual cells.
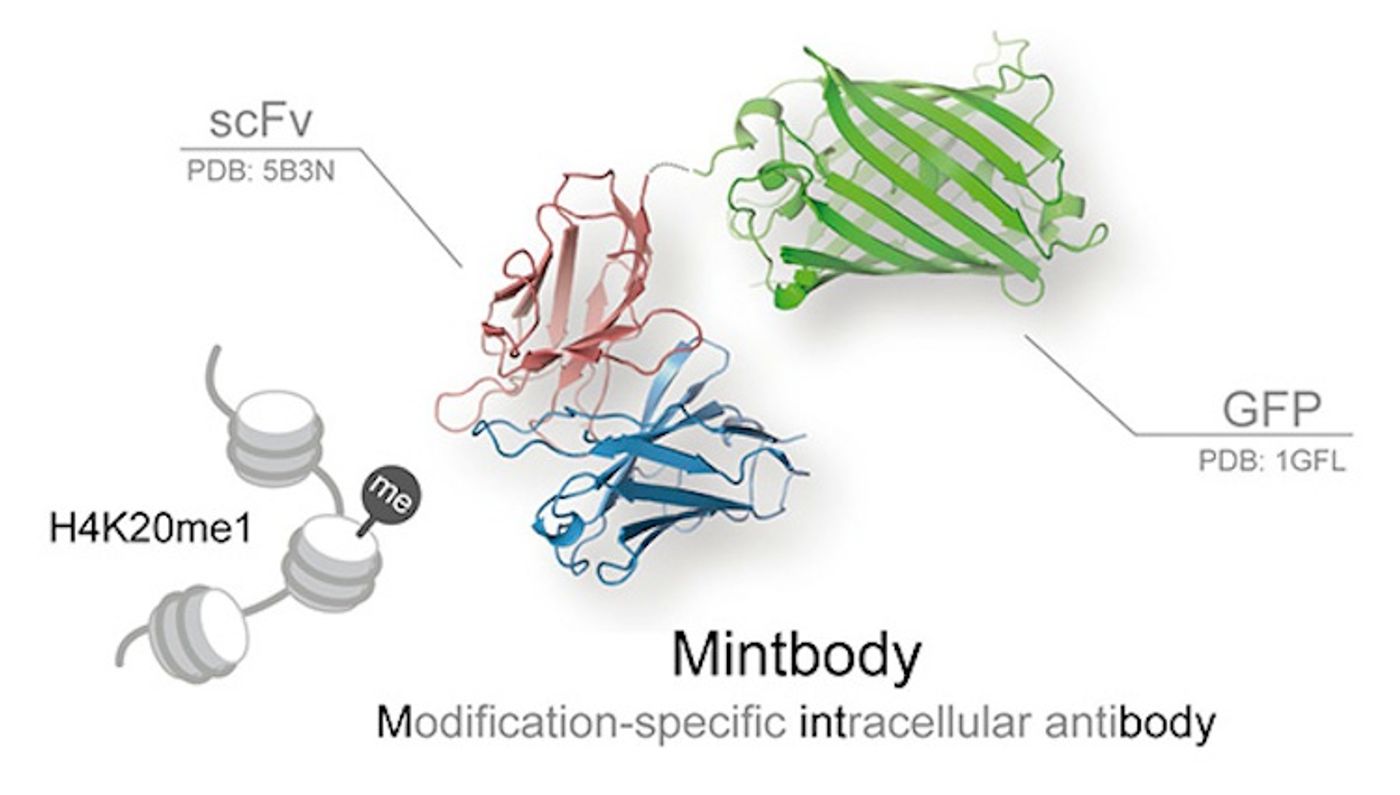
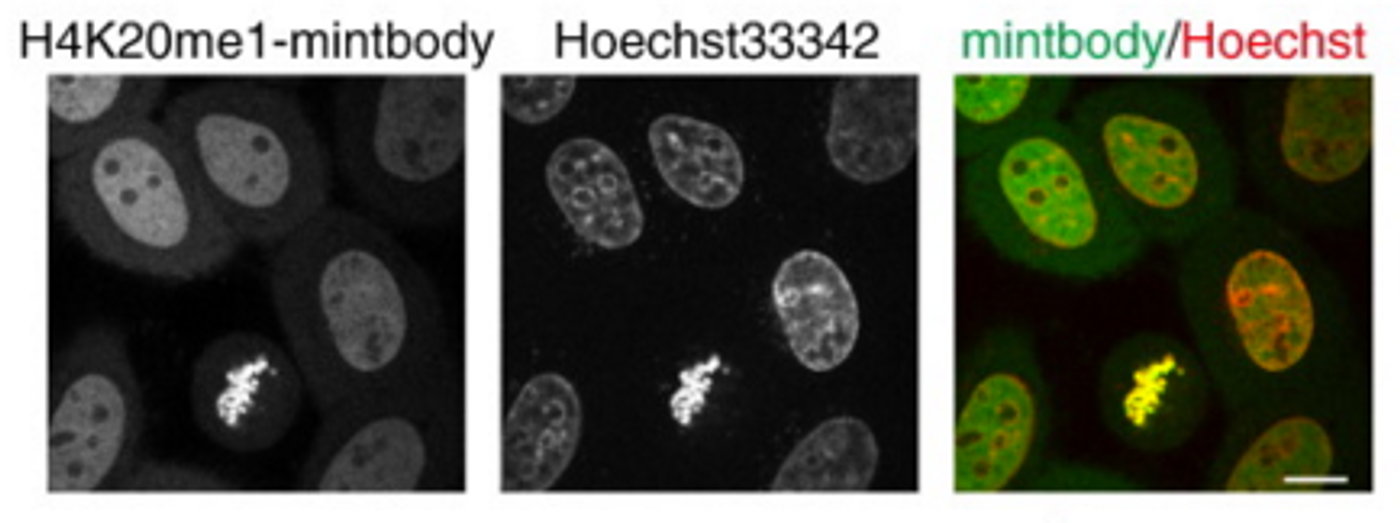
Publishing in the Journal of Molecular Biology, a research group led by Professor Kimura at the Institute of Innovative Research of the Tokyo Institute of Technology has developed a sensitive mintbody probe that will detect H4K20 monomethylation (H4K20me1) in living cells. Now, the intracellular action of the modification can be observed. The probe is a single-chain variable fragment, a fusion protein containing portions of immunoglobulins linked to a peptide, which has been fused to a fluorescent tag. The researchers indicate that it is very specific to only the monomethylated state.
The investigators used the roundworm Caenorhabditis elegans to demonstrate that the H4K20me1-mintbody is able to illustrate changes in H4K20me1 over course of the cell cycle as well as the cellular location of X-inactivated chromosomes, all without disrupting cellular functions.
Using genetic analysis and X-ray crystallography, the new work has also identified amino acids that are critical to the solubility and conformational stability of the H4K20me1-mintbody. Aberrant folding of the antibody fragments in the cellular cytoplasm usually causes solubility problems, but a potential solution has been found. As such, the researchers have overcome challenges to the performance of antibody fragments in live cells due to solubility issues.
The continued development of mintbodies for various post-translational modifications to histone opens up the door to identifying mechanism that underlie regulation of epigenetic changes.
Source:
Phys.org via Tokyo Institute of Technology,
Journal of Molecular Biology